curl examples. Note that curl is used for illustration only, any HTTP client will do. Alternatively, the /ocpu/test page can be used to explore the API.
HTTP in OpenCPU
The OpenCPU root path
The root of the API is dynamic. It defaults to /ocpu/, however system administrators can change this. Clients should make the OpenCPU server address and root path configurable. In the examples below we assume the default /ocpu/.
Debugging
The /ocpu/info endpoint shows the output of sessionInfo() from the main (webserver) process. This can be helpful for debugging. The /ocpu/test URL gives you a handy testing web page to perform server requests.
HTTP Methods
OpenCPU currently only uses HTTP methods GET and POST. GET is used to retrieve a resource, and POST is used for RPC. A POST request is only valid targeting a script or function URL.
| Method | Target | Action | Arguments | Example |
|---|---|---|---|---|
GET |
object | read object | control output format | GET /ocpu/library/MASS/R/cats/json |
POST |
object | call function | function arguments | POST /ocpu/library/stats/R/rnorm |
GET |
file | read file | - | GET /ocpu/library/MASS/NEWS GET /ocpu/library/MASS/scripts/ |
POST |
file | run script | control interpreter | POST /ocpu/library/MASS/scripts/ch01.R POST /ocpu/library/knitr/examples/knitr-minimal.Rmd |
#curl uses http get method by default
curl https://cloud.opencpu.org/ocpu/library/MASS/data/Boston/json
curl https://cloud.opencpu.org/ocpu/library/MASS/NEWS
curl https://cloud.opencpu.org/ocpu/library/MASS/scripts/
#curl uses http post method for -X POST or -d "arg=value"
curl https://cloud.opencpu.org/ocpu/library/MASS/scripts/ch01.R -X POST
curl https://cloud.opencpu.org/ocpu/library/stats/R/rnorm -d "n=10&mean=5"HTTP Status Codes
These are common statuscodes returned by OpenCPU that the client should be able to interpret
| HTTP Code | When | Returns |
|---|---|---|
200 OK |
On successful GET request | Resource content |
201 Created |
On successful POST request | Output location |
302 Found |
Redirect | Redirect Location |
400 Bad Request |
R raised an error. | Error message in text/plain |
502 Bad Gateway |
Nginx (opencpu-cache) can't connect to OpenCPU server. | (admin needs to look in error logs) |
503 Bad Request |
Serious problem with the server | (admin needs to look in error logs) |
API Endpoints
The API Libraries
The following libraries hold packages and sessions that are available on the server
| Path | What |
|---|---|
/ocpu/library/{pkgname}/ |
R packages installed in one of the global libraries on the server. |
/ocpu/apps/{gituser}/{reponame}/ |
Interfaces to R package installed in the root of repository {reponame} from github user {gituser}. |
/ocpu/user/{username}/library/{pkgname}/ |
R packages installed in the home library of Linux user {username}. |
/ocpu/tmp/{key}/ |
Temporary sessions, which hold outputs from a function/script RPC. |
#read packages
curl https://cloud.opencpu.org/ocpu/library/
curl https://cloud.opencpu.org/ocpu/apps/rwebapps/
curl https://cloud.opencpu.org/ocpu/user/jeroen/library/
#read session
curl https://cloud.opencpu.org/ocpu/tmp/x2c5ab8d4d6The R package API
Any of the above /{package}/ libraries support the following endpoints:
| Path | What |
|---|---|
../{pkgname}/info |
Show information about this package. |
../{pkgname}/www/ |
Apps (if any) included with the package |
../{pkgname}/R/ |
R objects exported by this package. See R object API. |
../{pkgname}/data/ |
Data included with this package. Datasets are objects, see R object API. |
../{pkgname}/man/ |
Manuals (help pages) included in this package. |
../{pkgname}/man/{topic}/{format} |
Retrieve help page about topic in output format format. Manual format must be one of text, html, pdf |
../{pkgname}/html |
Simulates the R-base html help pages (for backward compatibility). |
../{pkgname}/* |
For all else, interfaces to the files in the package installation directory. |
#package info
curl https://cloud.opencpu.org/ocpu/library/MASS/
#package objects (mostly functions)
curl https://cloud.opencpu.org/ocpu/library/MASS/R/
curl https://cloud.opencpu.org/ocpu/library/MASS/R/rlm/print
#package data objects
curl https://cloud.opencpu.org/ocpu/library/MASS/data/
curl https://cloud.opencpu.org/ocpu/library/MASS/data/housing/json
#read manuals pages
curl https://cloud.opencpu.org/ocpu/library/MASS/man/
curl https://cloud.opencpu.org/ocpu/library/MASS/man/rlm/text
curl https://cloud.opencpu.org/ocpu/library/MASS/man/rlm/html
curl https://cloud.opencpu.org/ocpu/library/MASS/man/rlm/pdf
#read files included with this package
curl https://cloud.opencpu.org/ocpu/library/MASS/scripts/
curl https://cloud.opencpu.org/ocpu/library/MASS/scripts/ch01.R
curl https://cloud.opencpu.org/ocpu/library/MASS/NEWS
#call a function (example from 'rlm' help page)
curl https://cloud.opencpu.org/ocpu/library/MASS/R/rlm -d "formula=stack.loss ~ .&data=stackloss&psi=psi.bisquare"
#run R script
curl https://cloud.opencpu.org/ocpu/library/MASS/scripts/ch01.R -X POST
#read output (replace key with value returned from previous request)
curl https://cloud.opencpu.org/ocpu/tmp/x0648ec526b/R/.val/printThe R object API
The /R API is used to read R objects, or call R functions.
| Path | What |
|---|---|
../R/ |
List R objects in this package or session. |
../data/ |
List data objects in a package. |
../{R|data}/{object} |
Read object in default format. If object is a function, it can be called using HTTP POST. |
../{R|data}/{object}/{format} |
Retrieve an R object in a particular output format (see section on output formats). |
#list objects and datasets from MASS
curl https://cloud.opencpu.org/ocpu/library/MASS/R/
curl https://cloud.opencpu.org/ocpu/library/MASS/data/
#retrieve objects
curl https://cloud.opencpu.org/ocpu/library/MASS/R/truehist/print
curl https://cloud.opencpu.org/ocpu/library/MASS/data/bacteria/print
curl https://cloud.opencpu.org/ocpu/library/MASS/data/bacteria/json
curl https://cloud.opencpu.org/ocpu/library/MASS/data/bacteria/csv
curl https://cloud.opencpu.org/ocpu/library/MASS/data/bacteria/rda
#call a function
curl https://cloud.opencpu.org/ocpu/library/MASS/R/truehist -d "data=[1,3,7,4,2,4,2,6,23,13,5,2]"The R session API
A session is a container that holds resources created from a remote function/script call (RPC).
| Path | What |
|---|---|
/ocpu/tmp/{key}/ |
List available output for this session. |
/ocpu/tmp/{key}/R |
R objects stored in this session. Interface using R object API, same as objects in packages. |
/ocpu/tmp/{key}/graphics/ |
Graphics (plots) stored in this session. |
/ocpu/tmp/{key}/graphics/{n}/{format} |
Retrieve plot number {n} in output format {format}. Format is usually one of png, pdf or svg. The {n} is an integer or "last". |
/ocpu/tmp/{key}/source |
Reads the input source code for this session. |
/ocpu/tmp/{key}/stdout |
Shows text printed to STDOUT in this session. |
/ocpu/tmp/{key}/console |
Shows the console input/output for this session (combines source and stdout) |
/ocpu/tmp/{key}/zip |
Download the entire session as a zip archive. |
/ocpu/tmp/{key}/tar |
Download the entire session as a gzipped tarball. |
/ocpu/tmp/{key}/files/* |
Interfaces to the file API in the working dir of the session. |
#A POST will create a temporary session
#Use the returned key for the subsequent calls below
curl https://cloud.opencpu.org/ocpu/library/MASS/scripts/ch01.R -X POST
#Look at console input/output
curl https://cloud.opencpu.org/ocpu/tmp/x05b85461/console/text
#We read session R objects
curl https://cloud.opencpu.org/ocpu/tmp/x05b85461/
curl https://cloud.opencpu.org/ocpu/tmp/x05b85461/R/
curl https://cloud.opencpu.org/ocpu/tmp/x05b85461/R/dd/csv
curl https://cloud.opencpu.org/ocpu/tmp/x05b85461/R/t.stat/print
#Or even call a function
curl https://cloud.opencpu.org/ocpu/tmp/x05b85461/R/t.stat -d "x=[1,0,0,1,1,1,0,1,1,0]"
#Download file from the working dir
curl https://cloud.opencpu.org/ocpu/tmp/x05b85461/files/ch01.pdf
#Check sessionInfo
curl https://cloud.opencpu.org/ocpu/tmp/x05b85461/info/printInput, Output: Data and Formats
Output formats for R objects
Any R object (including recorded graphics) can be retrieved in various output formats, indicated by :format in the API tables.
| Format | Content-type | Encoder (+args) | Data Frame | Matrix | List | Graphics | Other (function, S4) |
|---|---|---|---|---|---|---|---|
print |
text/plain | base::print | |||||
json |
application/json | jsonlite::toJSON | |||||
ndjson |
application/ndjson | jsonlite::stream_out | |||||
csv |
text/csv | utils::write.csv | |||||
tab |
text/plain | utils::write.table | |||||
md |
text/markdown | pander::pander | |||||
rda |
application/octet-stream | base::save | |||||
rds |
application/octet-stream | base::saveRDS | |||||
pb |
application/x-protobuf | protolite::serialize_pb | |||||
feather |
application/feather | feather::write_feather | |||||
png |
image/png | grDevices::png | |||||
pdf |
application/pdf | grDevices::pdf | |||||
svg |
image/svg+xml | grDevices::svg |
#read R objects from packages.
curl https://cloud.opencpu.org/ocpu/library/datasets/R/mtcars/json?digits=0
curl https://cloud.opencpu.org/ocpu/library/datasets/R/mtcars/csv
curl https://cloud.opencpu.org/ocpu/library/datasets/R/mtcars/tab?sep="|"
curl https://cloud.opencpu.org/ocpu/library/MASS/R/loglm/print#create a simple plot
curl https://cloud.opencpu.org/ocpu/library/graphics/R/plot -d "x=cars"
#replace session id with returned one
curl https://cloud.opencpu.org/ocpu/tmp/x0468b7ab/graphics/last/png
curl https://cloud.opencpu.org/ocpu/tmp/x0468b7ab/graphics/1/png?width=1000
curl https://cloud.opencpu.org/ocpu/tmp/x0468b7ab/graphics/last/svg
curl https://cloud.opencpu.org/ocpu/tmp/x0468b7ab/graphics/last/pdf?width=8Argument formats for R function calls (HTTP POST only)
When calling a function, we need to pass arguments. OpenCPU accepts the following types of arguments
| Type | What | Examples |
|---|---|---|
| Primitives | Numbers, strings, booleans | 3.1415, false, "mytitle" |
| JSON object | An array or object | {"foo" : [1,2,3], "bar" : {"name" : "jerry"}} |
| R code | plain R code | paste("useR", "2013"), mtcars |
| File upload | File argument will be temporary file path. | (multipart only) |
| Temp key (session) | Session ID from previous function call. | x2ac58d69 |
When performing a function call, the POST data (request body) can be formated using either multipart/form-data,
application/x-www-form-urlencoded, application/json or application/x-protobuf. Not every content-type supports any argument format:
| Content-type | Primitives | Data structures | R code | File upload | Temp key (session) |
|---|---|---|---|---|---|
application/x-www-form-urlencoded |
(inline json) | ||||
multipart/form-data |
(inline json) | ||||
application/json |
|||||
application/x-protobuf |
#call some functions
curl https://cloud.opencpu.org/ocpu/library/stats/R/rnorm -d "n=10&mean=5"
curl https://cloud.opencpu.org/ocpu/library/graphics/R/hist -d "x=[2,3,2,3,4,3,3]&breaks=10"
curl https://cloud.opencpu.org/ocpu/library/graphics/R/plot -d "x=cars&main='test'"
curl https://cloud.opencpu.org/ocpu/library/base/R/identity -d "x=coef(lm(speed~dist, data=cars))"#upload local file mydata.csv
curl https://cloud.opencpu.org/ocpu/library/utils/R/read.csv -F "file=@mydata.csv"
#replace session id with returned one above
curl https://cloud.opencpu.org/ocpu/tmp/x067b4172/R/.val/print
curl https://cloud.opencpu.org/ocpu/library/base/R/summary -d "object=x067b4172"#post arguments in json
curl http://cloud.opencpu.org/ocpu/library/stats/R/rnorm \
-H "Content-Type: application/json" -d '{"n":10, "mean": 10, "sd":10}'Running scripts and reproducible documents
We can run a script by performing HTTP POST on a file. The script is interpreted according to its (case insensitive) file extension. Any HTTP POST arguments are passed on to the interpreting function. The following types are supported:
| File extension | Type | Interpreter | Arguments |
|---|---|---|---|
file.r |
R script | evaluate::evaluate | - |
file.tex |
latex | tools::texi2pdf | - |
file.rnw |
knitr/sweave | knitr::knit + tools::texi2pdf | - |
file.md |
markdown | knitr::pandoc | format (see ?pandoc) |
file.rmd |
knitr/markdown | knitr::knit + knitr::pandoc | format (see ?pandoc) |
file.brew |
brew | brew::brew | output (see ?brew) |
#run scripts which exist in packages
curl https://cloud.opencpu.org/ocpu/library/MASS/scripts/ch01.R -X POST
curl https://cloud.opencpu.org/ocpu/library/brew/featurefull.brew -X POST
curl https://cloud.opencpu.org/ocpu/library/brew/brew-test-2.brew -d "output=output.html"
curl https://cloud.opencpu.org/ocpu/library/knitr/examples/knitr-minimal.Rmd -X POST
curl https://cloud.opencpu.org/ocpu/library/knitr/examples/knitr-minimal.Rmd -d "format=docx"
curl https://cloud.opencpu.org/ocpu/library/knitr/examples/knitr-minimal.Rmd -d "format=html"
curl https://cloud.opencpu.org/ocpu/library/knitr/examples/knitr-minimal.RnwJSON I/O RPC (a.k.a. Data Processing Units)
For the common special case where a client is only interested in the output data from a function call in JSON format, the HTTP POST request url can be post-fixed with /json. In this case, a successfull call will return status 200 (instead of 201), and the response body directly contains the returned object in JSON; no need for an additional GET request.
curl http://cloud.opencpu.org/ocpu/library/stats/R/rnorm/json -d n=2
[
-1.2804,
-0.75013
]We can combine this with the application/json request content type for full JSON RPC on R functions
curl http://cloud.opencpu.org/ocpu/library/stats/R/rnorm/json \
-H "Content-Type: application/json" -d '{"n":3, "mean": 10, "sd":10}'
[
4.9829,
6.3104,
11.411
]The above request invokes the following R function call:
library(jsonlite)
args <- fromJSON('{"n":3, "mean": 10, "sd":10}')
output <- do.call(stats::rnorm, args)
toJSON(output)Which in this case is equivalent to:
rnorm(n=3, mean=10, sd=10)
Miscellaneous functionality
OpenCPU Apps
OpenCPU apps are static web pages (html, css, js) which are included in an R package. They interface the R functions in this package through the OpenCPU API's. By convention, these apps are placed in the /inst/www/ directory of the R source package. See for more information the apps page.
Github CI Hook
The OpenCPU cloud server includes support for continuous integration (CI). Thereby a Github repository can be configured to automatically install your package on an OpenCPU server, every time a commit is pushed to the master branch. To take advantage of this feature, it is required that:
- The R source package is in the root directory of your repository. (example)
- The Github user account has a public email address
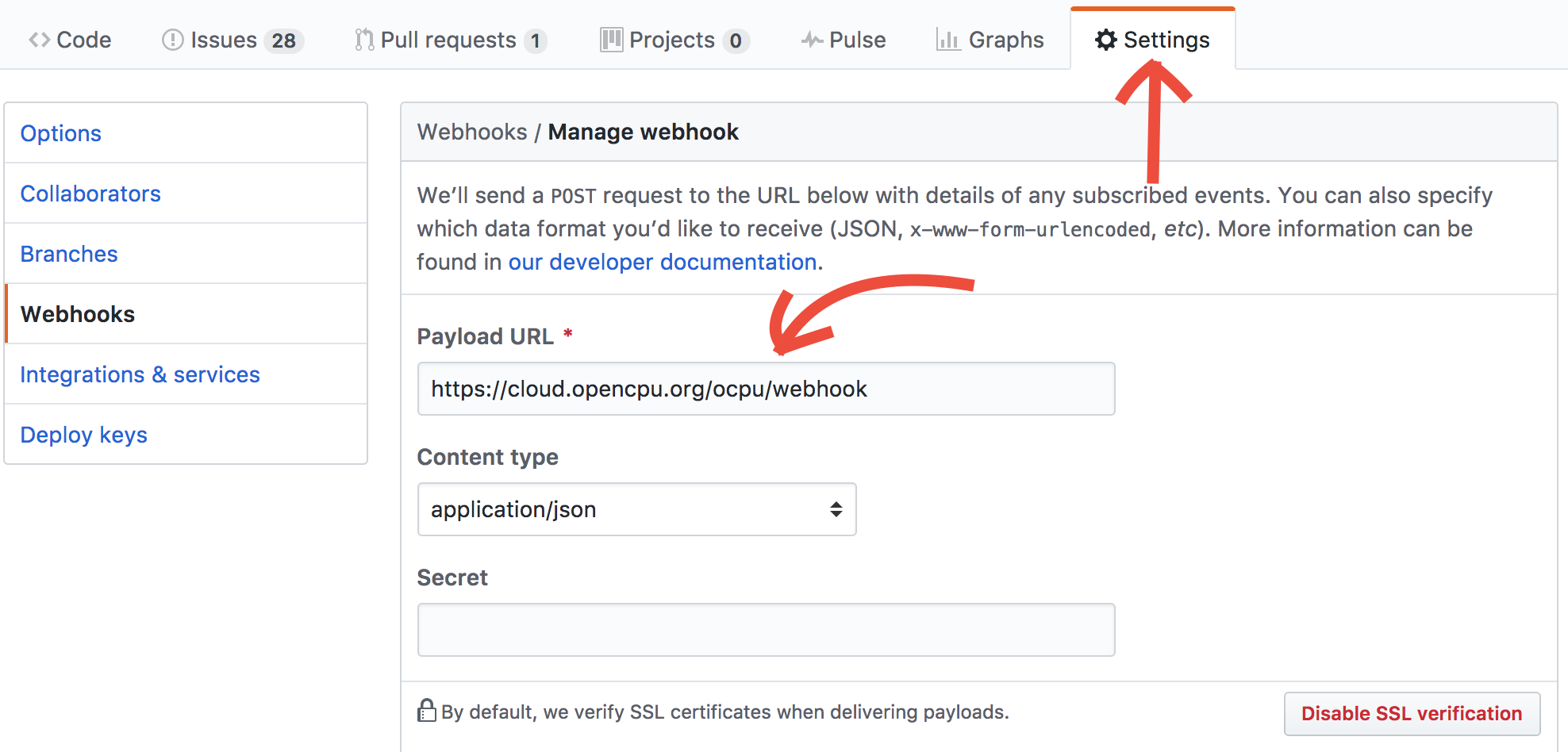
To setup CI, add the /ocpu/webhook URL for your server as a 'WebHook' in your Github repository. For example, to use the public demo server, add a webhook with the URL below (you can leave the Content-type and Secret field unchanged)
https://cloud.opencpu.org/ocpu/webhook To trigger a build, push a commit to the master branch. The build will show up under Recent Deliveries in your github webhook page, and you should receive an email (in your spam folder) reporting if the installation was successful. If it was, the package will directly be available for remote use at /ocpu/apps/{username}/{package}/ on the server.
If you are using the public server, the package will also be available via https://{yourname}.ocpu.io/{package}/. If you are running your own OpenCPU cloud server, an SMTP server must be configured for the email notifictions to work.
